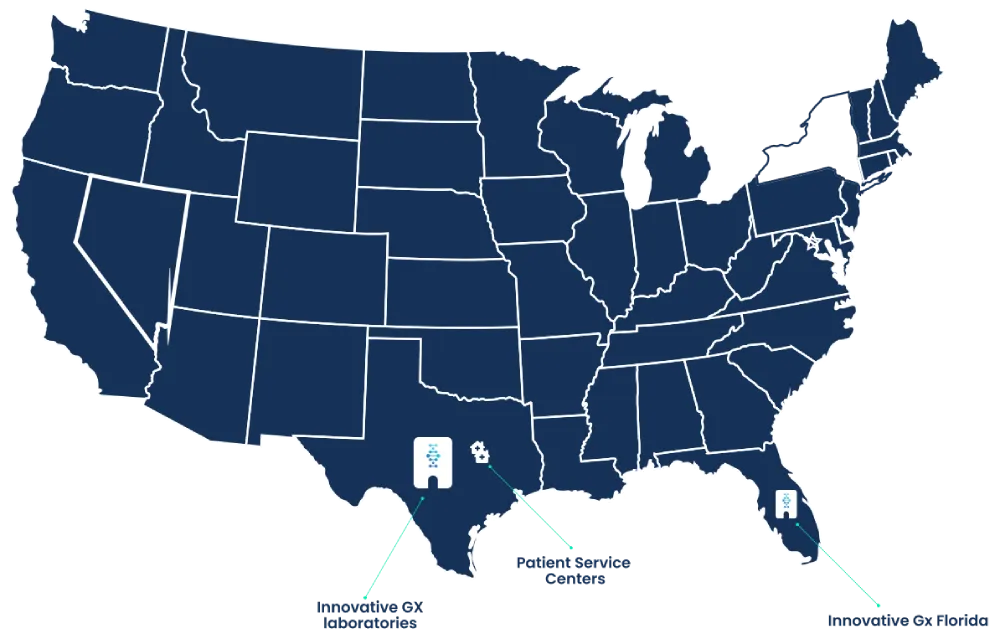
We have multiple clients in many areas that are integrated with through Athena.
Integrated with AdvancedMD
Integrated with Experity HealthCare
Supporting EMR’s: eClinical, Pratice Fusion, Quanum by Quest, Office Ally, PointClickCare, DrChrono, Kareo, Kipu, Medconnect, PrognoCIS, CharmEMR, NextGen Office, AdvancedMD.
Integrated with eClinicalWorks
IGx’s Quality Management System satisfies and surpasses industry quality standards and federal regulations.
Learn more